With the last release of our the Search & Match Service, the WMDA has maintained a similar haplotype frequency set configuration as was used in Optimatch (Search & Match v1). In most cases the ION of the donor/ cord blood unit will determine which haplotype frequency set is used.
To provide a more up-to-date representation of the donor pools world wide, the haplotype frequency set calculations have been performed using the latest data in the WMDA Donor and CBU database. Calcations were performed using the open source algorithm Haplo-o-Mat (https://github.com/DKMS/Hapl-o-Mat and https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5450239/) using high resolution typing results from organisations to extrapolate the haplotypes of the region or organisation. Thus, an organisation or geographical region must meet a minimum threshold of number of donors and availability of high resolution typing to build usable and valuable frequencies. For cord blood units. we will use the same sets that were determined in the donor populations.
The inclusion criteria aims to balance a high number of donors, the quality of the HLA types and the complexity of the haplotype frequency estimation. The number and HLA types of the loci that are included in the estimation can be independent of the loci of the haplotypes actually estimated if the quality of the former is appropriate.
5-locus Haplofrequency estimation was performed on all organisations/populations with at least 10000 donors in HLA-A, -B, -C, -DRB1 and -DQB1 types where ambiguity was low enough to be useful. Organisations in the same country/within the same population are usually combined, e.g. AT-ABMDR (ION-2614, Austrian Bone Marrow Donors) and AT-GFL (ION-4961, Verein Geben für Leben) but *not* for US-NMDP (general US population) and US-GOL (predominantly Ashkenazi Jews).
The current global frequency set will currently be used for all patients and for donors that for some reason do not match a typical haplotype frequency set. For example when a registry is new and has not yet been assigned to an appropriate existing haplotype frequency set or when the HLA typings from this registry deviate substantially enough from any typical haplotype frequency set that it does not make sense to assign them to one. The number of this frequency set is 999. We currently return the haplotype frequency set in the search results API endpoint for CBUs and will soon add it for donors.
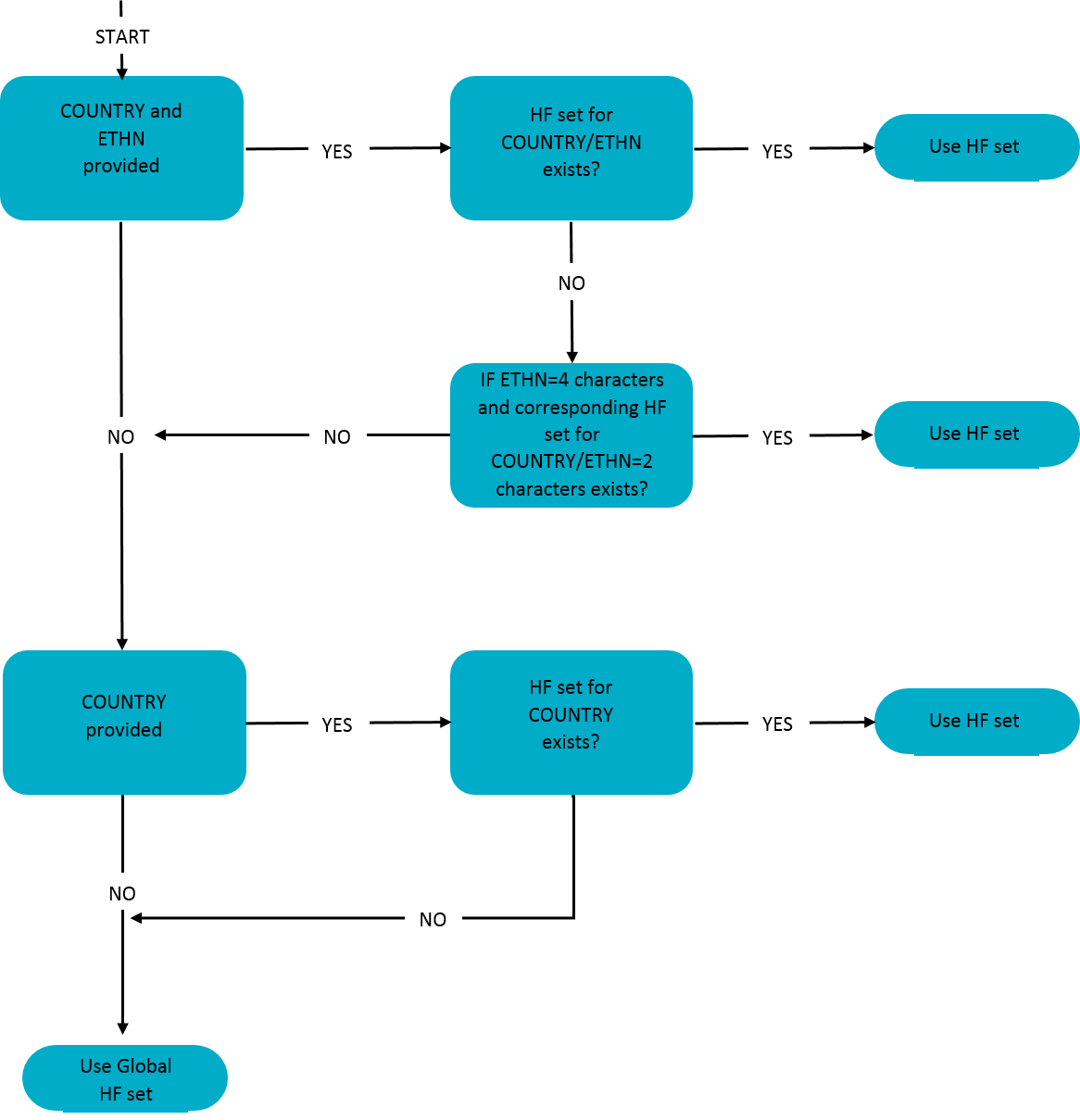
Below, you can find a figure how the haplotype frequency (HF) sets are assigned to the donors and cord blood units. The system first determines if both registry ION and ethnicity are provided. Registry ION is then used to look up the country/ region that it belongs to. If yes, then it will check if an ethnicity specific set for this registry exists. If the donor for example is coming from USA and has ethnicity HISA, then we do not have a specific set available. However, there is a more broad Hispanic HF set available (USA-HI) and this will then be used.
If there is no ethnicity available for the donor or no ethnicity specific HF set exists for that country, the system will check if a country specific HF set is available or if the country is part of one of the regional HF sets.
If no country specific HF set is available, the Search & Match Service will use the global consensus HF set.
Flowchart assigning a haplotype frequency (HF) set
In the table below, you can find the haplotype frequency set numbers and which countries/organisations were included as well as the total sample size. In the last part from the table we included regional sets, some country/ethnicity specific sets and the global consensus set.
The regional sets are defined as follows:
East Asia (eas): CN (CN+CN1), HK, TW, JP (Set number: 32)
Eastern Europe (eeu):AT (A+A2), CZ (CS+CS2), CY (CY+CY2), GR2, PL (PL3+PL5+PL6), TR (TRAN+TRIS+TRKK), BG, HU, HR, LT, MK, RU (R2+R4), RO, RS, SK, SI (Set number: 33)
South America (sam): AR, BR, UY (Set number: 34)
INPUT DATA | |||||
HF set number(pop_id) | Determination date | Sample size | ISO country code | ION | Search & Match Service_Registry_codes |
| 1 | 2022-05 | 226507 | AR | 5117 | AR-INCUCAI |
| 2 | 2022-05 | 149750 | AT | 2614 4961 | AT-ABMDR AT-Verein |
| 3 | 2022-05 | 41348 | AU + NZ | 7748 8261 | AU-ABMDR NZ-NZBMDR |
| 4 | 2022-05 | 36351 | BE | 4201 | BE-MDPB |
| 5 | 2022-05 | 81859 | BR | 8766 | BR-REDOME |
| 6 | 2022-05 | 68077 | CA | 5103 6912 | CA-One Match CA-HemaQuec |
| 7 | 2022-05 | 114900 | CH | 9341 | CH-SBSC |
| 8 | 2022-05 | 564150 | CN + HK + TW | 2197 6681 4070 3458 | CN-CMDP CN-Sunsh HK-HKBMDR TW-Tzu Chi |
| 9 | 2022-05 | 74633 | CY | 4278 9751 | CY-Par BMDR CY-CBMDR |
| 10 | 2022-05 | 75810 | CZ | 4753 5440 | CZ-CSCR CZ-CNMDR |
| 11 | 2022-05 | 2627544 | DE | 6939 | DE-ZKRD |
| 12 | 2022-05 | 53849 | DK | 2015 7484 | DK-DSCDW DK-DSDE |
| 13 | 2022-05 | 81265 | ES | 7813 | ES-REDMO |
| 14 | 2022-05 | 39567 | FI | 9738 | Fi-FSCR |
| 15 | 2022-05 | 90069 | FR | 1804 | FR-FGM |
| 16 | 2022-05 | 636595 | GB + IE | 6354 1726 2731 9968 5590 | GB-Anthony GB-WBMDR GB-BBMR GB-DKMS IE-IUBMR |
| 17 | 2022-05 | 102358 | GR | 1461 | GR-CBMDP |
| 18 | 2022-05 | 651085 | IL | 5239 4987 4068 | IL-Hadassah IL-Ezer Miz. IL-SHBB |
| 19 | 2022-05 | 530272 | IN | 8486 2824 4131 4596 | IN-Datri IN-GeneBand IN-MDR IN-BMST |
| 20 | 2022-05 | 84477 | IT | 7450 | IT-IBMDR |
| 21 | 2022-05 | 328557 | NL | 8139 | NL-Matchis |
| 22 | 2022-05 | 16499 | NO | 7214 | NO-NBMDR |
| 23 | 2022-05 | 1464544 | PL | 3918 5391 7414 | PL-ALF PL-Poltranspl PL-DKMS |
| 24 | 2022-05 | 11029 | PT | 7258 | PT-Cedace |
| 25 | 2022-05 | 97139 | SA | 8118 | SA-SSCDR |
| 26 | 2022-05 | 184283 | SE | 5285 | SE-Tobias |
| 27 | 2022-05 | 78885 | SG | 3785 | SG-BMDP |
| 28 | 2022-05 | 122133 | TH | 8362 | TH-TSCDR |
| 29 | 2022-05 | 491941 | TR | 3893 5509 3503 | TR-TRAN TR-TRIS TR-TURKOK |
| 30 | 2022-05 | 1723292 | US | 3553 | US-NMDP |
| 31 | 2022-05 | 97953 | US | 1033 | US-GOL |
| 32 | 2022-05 | 564561 | CN+HK+TW+JP | eas | Search&Match:eas |
| 33 | 2022-05 | 2398213 | AT+CZ+CY+GR+PL+TR+BG+HU+ HR+LT+MK+RU+RO+RS+SK+SI | eeu | Search&Match:eeu |
| 34 | 2022-05 | 413438 | AR+BR+UY | sam | Search&Match:sam |
| 100 | 2022-05 | 151,204 | DE-ASSW | - | DE-ASSW subset |
| 101 | 2022-05 | 656,591 | US-AF (ZKRD) | - | US-AF subset (ZKRD_estimation) |
| 102 | 2022-05 | 796,780 | US-AS (ZKRD) | - | US-AS subset (ZKRD_estimation) |
| 103 | 2022-05 | 3,740,668 | US-CA (ZKRD) | - | US-CA subset (ZKRD_estimation) |
| 104 | 2022-05 | 1,002,893 | US-HI (ZKRD) | - | US-HI subset (ZKRD_estimation) |
| 999 | 2022-05 | 11,430,561 | Global consensus | ||
Since WMDA is doing these calculations on behalf of organisations, it is important for organisations to submit high resolution typing when available and donor ethnicity data. This greatly improves the haplofrequency sets generated and allows for individual donor match grades to be more accurate.
It is important to know that your organisations frequency set is changeable!
You can request that your donors be returned to the global consensus haplofrequency set or to a regional set. Your request should include the reasons why you believe that another set is more applicable for your donors. Additionally, if your organisation has a frequency set available that you would like to see utilized for the donors of your organisation we can implement it as well. In this case, your request should include at least a description about your population, the sample size, inclusion and exclusion criteria, calculation method, and the reasons why your haplofrequency set would be better than the current set applied by WMDA. Both requests will be reviewed by the WMDA BioInformatics Working Group. Please send requests to support@wmda.info . In the future, you'll be able to dictate a frequency set on a per donor basis.
| Date | Version | Description | Author |
|---|---|---|---|
| 2017-10-31 | 1.0 | Replacement BMDW global haplotype frequency set for more specific sets | JK |
| 2018-01-24 | 1.1 | Modification some sets; introduced during OptiMatch version 3.31.0 | JK |
| 2019-02-15 | 1.2 | Replaced BMDW by WMDA / Search & Match Service; updated email address | JK |