...
| Expand | ||
|---|---|---|
| ||
This can have several causes.
See the following page for more information: Feature differences Hap-E Search vs Optimatch |
| Expand | ||
|---|---|---|
| ||
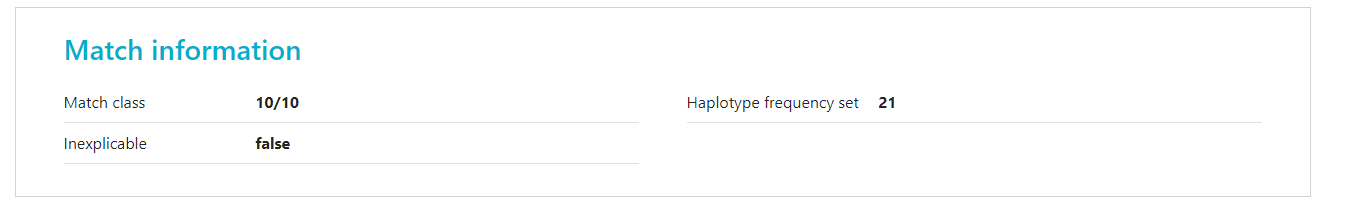
At the top of search results you can see how many donors or CBUs are inexplicable. Inexplicable donors or CBUs are records that have HLA that cannot be explained by a combination of two haplotypes in the haplotype frequency set (Haplotype frequency sets in the HAP-E matching algorithm of the Search & Match Service) for the population that the donor or CBU is part of. To find out which haplotype frequency set was used, you can use the full report The number there corresponds to the set on the Haplotype frequency sets in the HAP-E matching algorithm of the Search & Match Service share page. Because the HLA typing that this donor/CBU has cannot be explained, Hap-E also cannot calculate match probabilities. Currently these donors appear on the bottom of the match class that they are part of (e.g. 10/10, 9/10, 7/8). We are working on a way to move potentially relevant but inexplicable donors to a better place in the search results. The number of inexplicable donors/CBUs mentioned therefore serves as a reminder that there are potentially relevant search results available that can be found at the bottom of the match class. |
| Expand | ||
|---|---|---|
| ||
Since B*15:BPXE = B*15:03/61/74/103 a donor with this codes is a potential allele match for the patient. According to the official WMDA serology/DNA correspondence table, B*15:61 and B*15:74 have a serology of B15/B70 while B*15:03 is B72 and B*15:103 is B70. As a consequence a serology of B15 rules out B*15:03 and this donor is no longer potentially identical (on the allele level). Another explanation could be the limited length of donor lists. Background: Unfortunately, for many donors serology was derived from DNA (by using the first field for the serological assignment) and vice versa (by appending “:XX” to the serological assignment) and often eventually both values are reported. In the case discussed, B*15:BPXE probably was translated back into B15 which is most likely wrong. This is a typical and (with certain registries) frequent case but there are many more unexpected DNA-serology correspondences that can give rise to exactly the same situation. |
...